#include <Rule.h>

Public Member Functions | |
| Rule () | |
| Default constructor. More... | |
| virtual | ~Rule () |
| Default destructor. More... | |
| virtual Rule * | objFactory ()=0 |
| virtual Rule * | clone () |
| virtual void | copy (Rule *fromRule) |
| void | RestoreRule (double *perf, unsigned char *genes, int arry_len, int *gene_index) |
| Restore Model. More... | |
| virtual void | log () |
| virtual char | type () const |
| virtual char * | toString () |
| virtual void | fromString (char *strRule)=0 |
| virtual char * | toXML () |
| virtual void | initialize (EnvCellSet *objEnvCellSet, const RuleSet *objRuleSet, bool *geneIsActivePtr, int *geneIndexPtr, int iActGenes)=0 |
| virtual bool | applyToCell (EnvCell *cell)=0 |
| virtual double | getCertainty (EnvCell *cell) |
| virtual double | getError (BYTE pred, EnvCell *cell) |
| virtual double | getStrength (EnvCell *cell)=0 |
| virtual bool | similar (Rule *objOtherRule) |
| virtual void | mutate (int intTemperature) |
| double | testWithData (EnvCellSet *objTrainSet) |
| bool | needsEvaluation () |
Protected Attributes | |
| BYTE * | Gene |
| BYTE vector containing the genes (representation of the variables in a Genetic Algorithm. More... | |
| int | intGenes |
| Number of genes stored by the rule. More... | |
| double | dblPerformance [10] |
| Vector for storing the performance values for the rule. More... | |
| bool | blnNeedsEvaluation |
| int | intGens |
| int | intTrials |
| int | intScreener |
| int | intScreen |
| int | intLength |
| int | intNumber |
| int | intConclusion |
| char | chrOrigin |
| char | chrPad |
| int | lId |
| int | iOrigGen |
| double | _pXYs |
| int | _no |
| double | _dA |
| double | _dSig |
| bool * | bGeneIsActive |
| int * | iGeneIndex |
| int | iActiveGenes |
Friends | |
| class | RuleSet |
| class | GarpAlgorithm |
| class | CJobResultValidator |
Detailed Description
Constructor & Destructor Documentation
| Rule::Rule | ( | ) |
Default constructor.
Definition at line 33 of file Rule.cpp.
References _dA, _dSig, _no, _pXYs, bGeneIsActive, blnNeedsEvaluation, chrOrigin, chrPad, dblPerformance, Gene, iActiveGenes, iGeneIndex, intConclusion, intGenes, intGens, intLength, intNumber, intScreen, intScreener, intTrials, iOrigGen, and lId.
|
virtual |
Member Function Documentation
|
pure virtual |
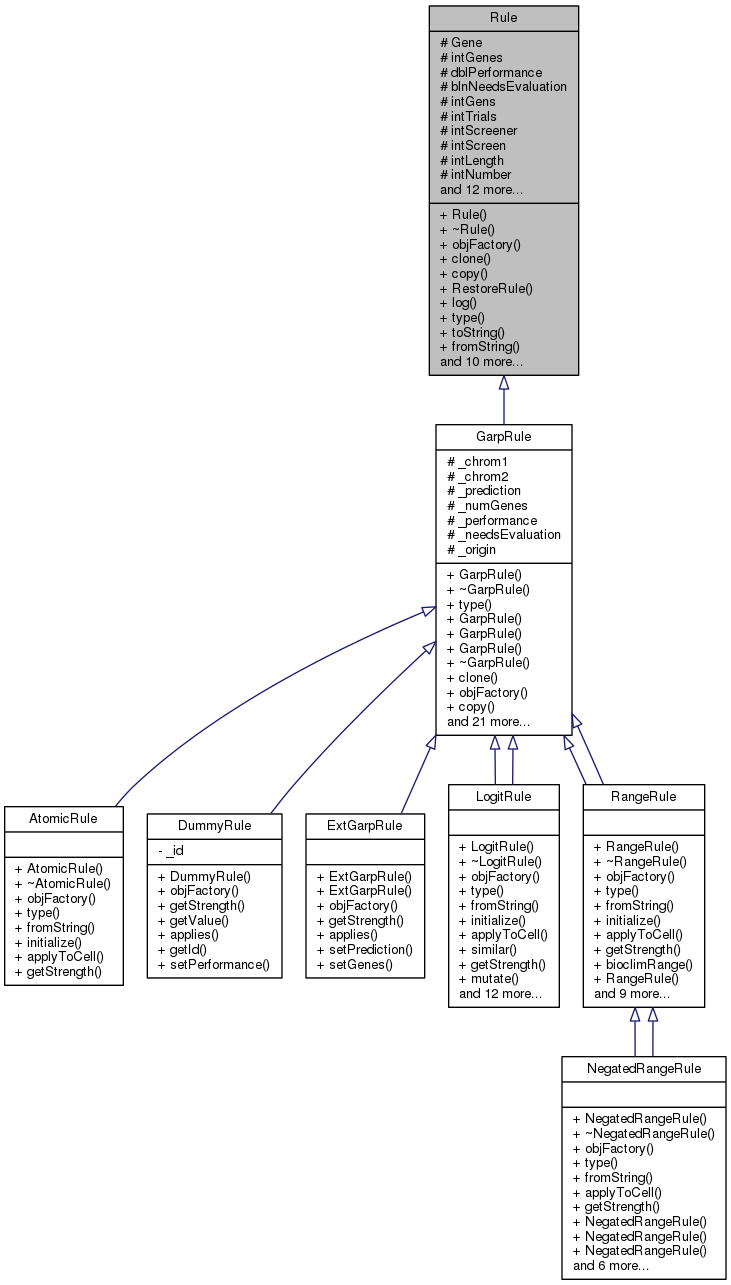
Implemented in LogitRule, AtomicRule, NegatedRangeRule, and RangeRule.
|
virtual |
Definition at line 73 of file Rule.cpp.
References _dA, _dSig, _no, _pXYs, bGeneIsActive, blnNeedsEvaluation, chrOrigin, chrPad, dblPerformance, Gene, iActiveGenes, iGeneIndex, intGenes, intGens, intLength, intScreen, intScreener, intTrials, iOrigGen, lId, and objFactory().
Referenced by GarpAlgorithm::concatenateRuleSets(), GarpAlgorithm::saveRule(), and GarpAlgorithm::select().

|
virtual |
Definition at line 115 of file Rule.cpp.
References _dA, _dSig, _no, _pXYs, bGeneIsActive, blnNeedsEvaluation, chrOrigin, chrPad, dblPerformance, Gene, iActiveGenes, iGeneIndex, intGenes, intGens, intLength, intScreen, intScreener, intTrials, iOrigGen, lId, and type().
Referenced by GarpAlgorithm::concatenateRuleSets().

|
pure virtual |
Implemented in LogitRule, AtomicRule, NegatedRangeRule, and RangeRule.
|
virtual |
Definition at line 305 of file Rule.cpp.
References Gene, and EnvCell::values.
Referenced by testWithData().
|
pure virtual |
Implemented in LogitRule, AtomicRule, NegatedRangeRule, and RangeRule.
Referenced by testWithData().
|
pure virtual |
Implemented in LogitRule, AtomicRule, and RangeRule.
Definition at line 180 of file Rule.cpp.
References bGeneIsActive, blnNeedsEvaluation, chrOrigin, chrPad, EnvCellSet::count(), Gene, EnvCellSet::get(), iActiveGenes, iGeneIndex, intConclusion, intGenes, intGens, intLength, intScreen, intTrials, GarpUtil::randint(), EnvCell::size(), type(), and EnvCell::values.
Referenced by GarpAlgorithm::colonize(), RangeRule::initialize(), AtomicRule::initialize(), and LogitRule::initialize().

|
virtual |
Reimplemented in GarpRule, RangeRule, NegatedRangeRule, and LogitRule.
Definition at line 904 of file Rule.cpp.
References dblPerformance, Gene, intGenes, and type().
Referenced by RuleSet::log().

|
virtual |
Reimplemented in LogitRule.
Definition at line 313 of file Rule.cpp.
References blnNeedsEvaluation, chrOrigin, Gene, iActiveGenes, iGeneIndex, intGens, intLength, GarpUtil::randint(), and type().
Referenced by GarpAlgorithm::mutate().

|
inline |
Definition at line 108 of file Rule.h.
References blnNeedsEvaluation.
Referenced by GarpAlgorithm::evaluate().
|
pure virtual |
Implemented in LogitRule, AtomicRule, NegatedRangeRule, and RangeRule.
Referenced by clone().
| void Rule::RestoreRule | ( | double * | perf, |
| unsigned char * | genes, | ||
| int | arry_len, | ||
| int * | gene_index | ||
| ) |
Restore Model.
Definition at line 156 of file Rule.cpp.
References dblPerformance, Gene, iActiveGenes, iGeneIndex, and intLength.
Referenced by GarpAlgorithm::_setConfiguration().
|
virtual |
Reimplemented in LogitRule.
Definition at line 349 of file Rule.cpp.
References Gene, iActiveGenes, iGeneIndex, intLength, and type().
Referenced by GarpAlgorithm::concatenateRuleSets(), and GarpAlgorithm::saveRule().

| double Rule::testWithData | ( | EnvCellSet * | objTrainSet | ) |
Definition at line 387 of file Rule.cpp.
References _dA, _dSig, _no, _pXYs, dblPerformance, EnvCellSet::get(), getCertainty(), getError(), getStrength(), MIN_SIG_NO, GarpUtil::randint(), and EnvCellSet::size().
Referenced by GarpAlgorithm::evaluate().

|
virtual |
|
virtual |
Definition at line 254 of file Rule.cpp.
References dblPerformance, Gene, intLength, iOrigGen, lId, and type().
Referenced by RuleSet::toXML().

|
inlinevirtual |
Reimplemented in LogitRule, AtomicRule, NegatedRangeRule, GarpRule, RangeRule, GarpRule, RangeRule, NegatedRangeRule, and LogitRule.
Definition at line 88 of file Rule.h.
Referenced by GarpAlgorithm::_getConfiguration(), GarpAlgorithm::concatenateRuleSets(), copy(), RuleSet::gatherRuleSetStats(), initialize(), log(), mutate(), GarpAlgorithm::saveRule(), similar(), toString(), and toXML().
Friends And Related Function Documentation
|
friend |
Member Data Documentation
|
protected |
|
protected |
|
protected |
|
protected |
|
protected |
Definition at line 71 of file Rule.h.
Referenced by clone(), copy(), initialize(), Rule(), and RuleSet::setActiveGenes().
|
protected |
Definition at line 52 of file Rule.h.
Referenced by clone(), copy(), GarpAlgorithm::crossover(), GarpAlgorithm::evaluate(), initialize(), GarpAlgorithm::join(), mutate(), LogitRule::mutate(), needsEvaluation(), Rule(), and GarpAlgorithm::select().
|
protected |
Definition at line 60 of file Rule.h.
Referenced by clone(), copy(), GarpAlgorithm::crossover(), initialize(), GarpAlgorithm::join(), mutate(), LogitRule::mutate(), Rule(), and GarpAlgorithm::saveRule().
|
protected |
|
protected |
Vector for storing the performance values for the rule.
Definition at line 51 of file Rule.h.
Referenced by GarpAlgorithm::_getConfiguration(), RuleSet::applyRulesToCell(), clone(), copy(), RuleSet::gatherRuleSetStats(), RuleSet::getOveralPerformance(), log(), GarpAlgorithm::measure(), RestoreRule(), Rule(), GarpAlgorithm::saveRule(), GarpAlgorithm::select(), testWithData(), toString(), and toXML().
|
protected |
BYTE vector containing the genes (representation of the variables in a Genetic Algorithm.
Definition at line 46 of file Rule.h.
Referenced by GarpAlgorithm::_getConfiguration(), RuleSet::applyRulesToCell(), RangeRule::applyToCell(), NegatedRangeRule::applyToCell(), AtomicRule::applyToCell(), RangeRule::bioclimRange(), clone(), copy(), GarpAlgorithm::crossover(), RuleSet::gatherRuleSetStats(), getCertainty(), RangeRule::getStrength(), AtomicRule::getStrength(), LogitRule::getStrength(), RuleSet::getValue(), initialize(), RangeRule::initialize(), AtomicRule::initialize(), LogitRule::initialize(), GarpAlgorithm::join(), log(), mutate(), LogitRule::mutate(), RestoreRule(), Rule(), similar(), LogitRule::similar(), toString(), toXML(), RuleSet::verify(), and ~Rule().
|
protected |
Definition at line 73 of file Rule.h.
Referenced by RangeRule::applyToCell(), NegatedRangeRule::applyToCell(), AtomicRule::applyToCell(), clone(), copy(), RangeRule::getStrength(), AtomicRule::getStrength(), LogitRule::getStrength(), initialize(), RangeRule::initialize(), AtomicRule::initialize(), LogitRule::initialize(), mutate(), LogitRule::mutate(), RestoreRule(), Rule(), RuleSet::setActiveGenes(), and similar().
|
protected |
Definition at line 72 of file Rule.h.
Referenced by RangeRule::applyToCell(), NegatedRangeRule::applyToCell(), AtomicRule::applyToCell(), clone(), copy(), RangeRule::getStrength(), AtomicRule::getStrength(), LogitRule::getStrength(), initialize(), RangeRule::initialize(), AtomicRule::initialize(), LogitRule::initialize(), mutate(), LogitRule::mutate(), RestoreRule(), Rule(), RuleSet::setActiveGenes(), and similar().
|
protected |
Definition at line 59 of file Rule.h.
Referenced by initialize(), and Rule().
|
protected |
Number of genes stored by the rule.
Definition at line 48 of file Rule.h.
Referenced by clone(), copy(), initialize(), log(), Rule(), and GarpAlgorithm::saveRule().
|
protected |
Definition at line 53 of file Rule.h.
Referenced by clone(), copy(), GarpAlgorithm::crossover(), GarpAlgorithm::evaluate(), initialize(), GarpAlgorithm::join(), mutate(), LogitRule::mutate(), and Rule().
|
protected |
Definition at line 57 of file Rule.h.
Referenced by GarpAlgorithm::_getConfiguration(), NegatedRangeRule::applyToCell(), clone(), copy(), GarpAlgorithm::crossover(), RangeRule::getStrength(), LogitRule::getStrength(), initialize(), RangeRule::initialize(), AtomicRule::initialize(), GarpAlgorithm::join(), mutate(), LogitRule::mutate(), RestoreRule(), Rule(), similar(), LogitRule::similar(), toString(), and toXML().
|
protected |
|
protected |
|
protected |
Definition at line 54 of file Rule.h.
Referenced by clone(), copy(), GarpAlgorithm::evaluate(), initialize(), Rule(), and GarpAlgorithm::saveRule().
|
protected |
|
protected |
The documentation for this class was generated from the following files:

